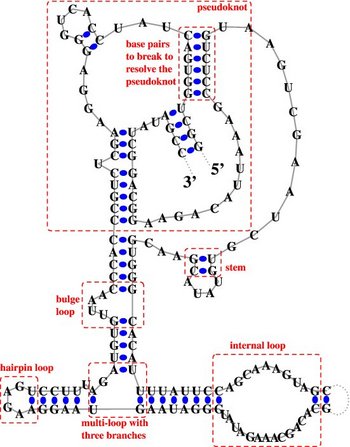
RNA STRAND v2.0 - The RNA secondary STRucture and statistical ANalysis Database
RNA STRAND contains known RNA
secondary structures of any type and organism. The
ultimate goal of this database is to incorporate a comprehensive
collection
of known RNA secondary structures, and to provide the scientific community
with
simple yet powerful ways of analysing, searching and updating the
proposed
database.
|
|||
|
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Publication
M. Andronescu, V. Bereg, H. H. Hoos, and A. Condon. RNA STRAND: The RNA Secondary Structure And Statistical Analysis Database. BMC Bioinformatics. 2008;9(1):340. [pubmed]
Supplementary material 1: List of Rfam published families which are included in RNA STRAND
Supplementary material 2: Description of the output of the RNA Secondary Structure Analyser
Download
The RNA Secondary Structure Analyser
The RNA STRAND database
Conditions of use
We hope that you find RNA STRAND useful. If you use the database or analysis provided on this website in your research, we ask you to acknowledge this by citing the [STRAND] reference below. We thank the following sources for giving us permission to redistribute RNA sequence and secondary structure data:
- RCSB Protein Data Bank: http://www.rcsb.org/pdb/
- Gutell Lab CRW Site: http://www.rna.ccbb.utexas.edu/
- tmRNA Database: http://rnp.uthct.edu/rnp/tmRDB/tmRDB.html
- Sprinzl tRNA Database: http://www.uni-bayreuth.de/departments/biochemie/sprinzl/trna/
- RNase P Database: http://jwbrown.mbio.ncsu.edu/RNaseP/home.html
- SRP Database: http://rnp.uthct.edu/rnp/SRPDB/SRPDB.html
- Rfam Database: http://www.sanger.ac.uk/Software/Rfam/index.shtml
- Nucleic Acid Database: http://ndbserver.rutgers.edu/
- [STRAND] Andronescu M, Bereg V, Hoos HH, and Condon A: RNA STRAND: The RNA Secondary Structure and Statistical Analysis Database. BMC Bioinformatics. 2008;9(1):340.
- [PDB] Westbrook J, Feng Z, Chen L, Yang H, Berman H: The Protein Data Bank and structural genomics. Nucleic Acids Res 2003, 31:489-491.
- [CRW] Cannone J, Subramanian S, Schnare M, Collett J, D'Souza L, Du Y, Feng B, Lin N, Madabusi L, Muller K, Pande N, Shang Z, Yu N, Gutell R: The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics 2002, 3:2. [Correction: BMC Bioinformatics 3:15.]
- [tmRDB,SRPDB] Andersen ES, Rosenblad MA, Larsen N, Westergaard JC, Burks J, Wower IK, Wower J, Gorodkin J, Samuelsson T, Zwieb C: The tmRDB and SRPDB resources. Nucleic Acids Res 2006, 34(Database issue):163-168.
- [SprinzlDB] Sprinzl M, Vassilenko K: Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res 2005, 33(Database issue):139-140.
- [RNasePDB] Brown J: The Ribonuclease P Database. Nucleic Acids Res 1999, 27:314-314.
- [Rfam] Griffiths-Jones S, Moxon S, Marshall M, Khanna A, Eddy S, Bateman A: Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res 2005, 33(Database issue):121-124.
- [NDB] Berman HM, Olson WK, Beveridge DL, Westbrook J, Gelbin A, Demeny T, Hsieh SH, Srinivasan AR, Schneider B: The nucleic acid database. A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys J 1992, 63(3):751-759.
Disclaimer
Use of the RNA STRAND is free of charge. Although the authors have made every effort to ensure that the database and analyzer are correctly implemented, and fulfill the function described in the documentation, neither the authors nor the University of British Columbia guarantee its correctness, fitness for a particular purpose, or future availability.
![]()
Copyright © 2004-2008 BETA LAB - University of British Columbia